Gene editing has undergone a revolution over the last decade, with CRISPR technologies moving from proof-of-concept experiments into early therapeutic applications. Among these, base editing and prime editing have emerged as two of the most promising approaches for correcting point mutations and small genetic defects without introducing double-strand breaks (DSBs). In 2025, both methods are actively evaluated for treating monogenic diseases, but they differ in mechanism, precision, efficiency, and clinical applicability.
This article provides a comparative overview of base editing and prime editing, highlighting their therapeutic potential and limitations.
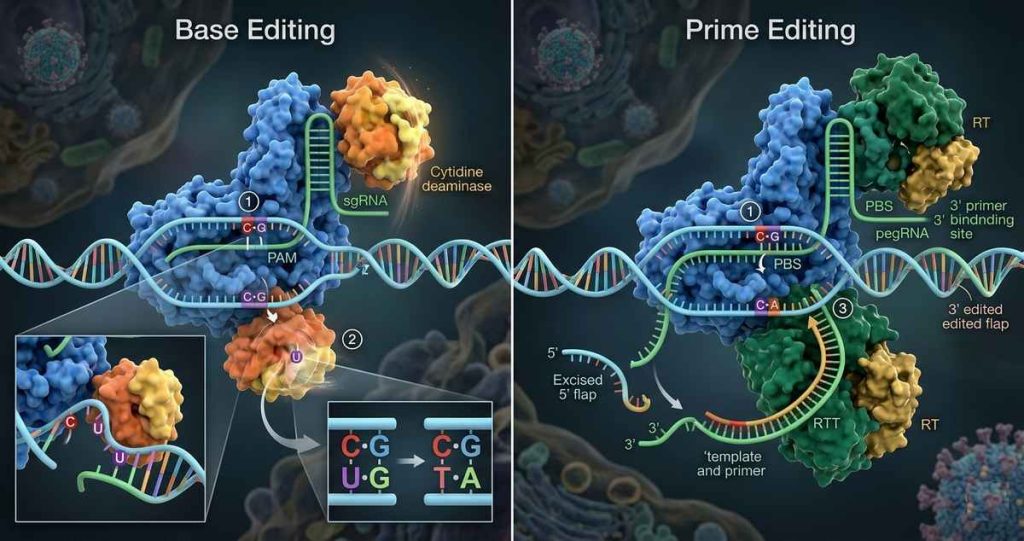
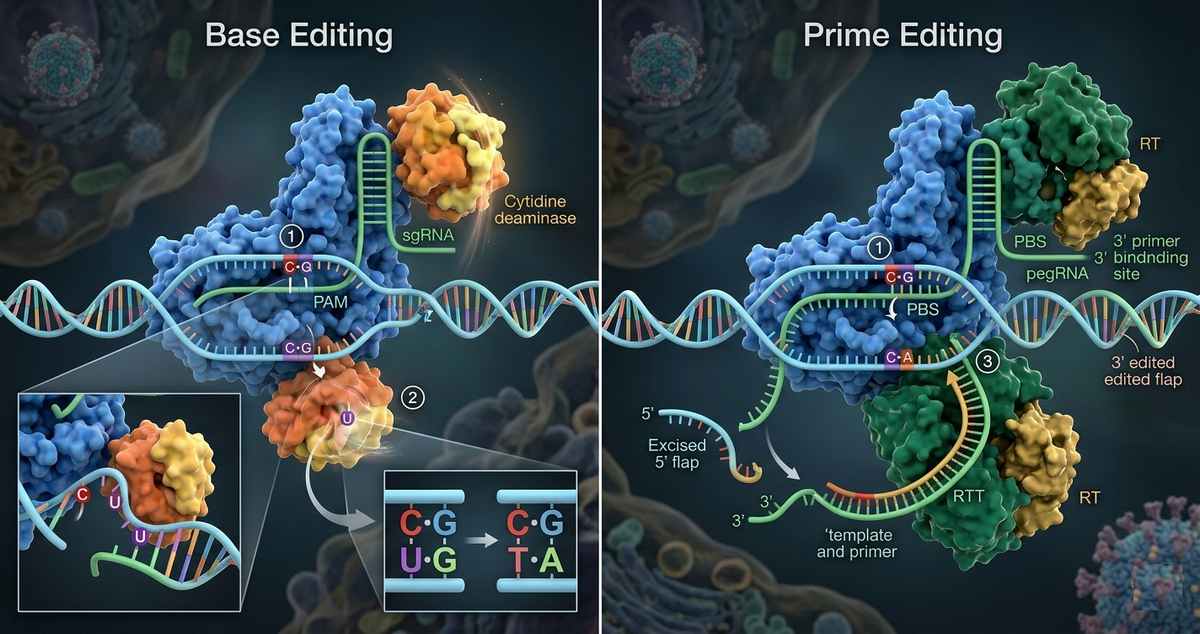
Mechanistic Overview
Base Editing
- Function: converts one nucleotide to another without cutting both DNA strands
- Mechanism: utilizes a catalytically impaired Cas protein fused to a deaminase enzyme
- Applications: single nucleotide polymorphism correction, in vivo gene therapy
- Advantages: high precision for specific base conversions (C→T, A→G)
- Limitations: restricted to certain nucleotide changes, potential off-target deamination
Prime Editing
- Function: programmable DNA writing tool capable of insertions, deletions, and all 12 base-to-base conversions
- Mechanism: uses Cas9 nickase fused to reverse transcriptase guided by a prime editing guide RNA (pegRNA)
- Applications: broader mutation correction, complex therapeutic edits
- Advantages: flexible, can correct multiple mutation types
- Limitations: lower efficiency in some cell types, pegRNA design complexity
Therapeutic Efficiency
| Feature | Base Editing | Prime Editing |
|---|---|---|
| Single-base conversion accuracy | High (90–99% in optimized systems) | Moderate to high (70–90%) |
| Editing window | Narrow (specific nucleotides) | Flexible, programmable |
| Targetable mutations | Limited (transition mutations) | Broad (transition + transversion, small indels) |
| Off-target risk | Low to moderate | Low but pegRNA-dependent |
| Delivery constraints | Smaller Cas-base editor fits in AAV | Larger, delivery more challenging |
Base editing often offers higher efficiency for targeted point mutations, making it attractive for diseases with single-base pathogenic variants. Prime editing, by contrast, is more versatile, addressing a wider range of mutation types but may require optimization for efficiency and delivery.
The future of precision medicine may also depend on computational modeling such as
digital twin patient systems that simulate treatment outcomes before clinical implementation.
Clinical Applications
Base Editing in 2025
- Hemoglobinopathies: sickle cell and beta-thalassemia
- Genetic retinal disorders
- Liver enzyme deficiencies
- Early-stage in vivo gene therapy trials
Prime Editing in 2025
- Correction of multi-base or complex point mutations
- Rare monogenic diseases with insertions or deletions
- Research pipelines for neuromuscular disorders
- Potential for ex vivo and in vivo therapeutic applications
Delivery and Safety Considerations
- Vector Compatibility
- Base editors are smaller and can fit within adeno-associated virus (AAV) vectors.
- Prime editors often require larger viral vectors or non-viral delivery systems.
- Immunogenicity
- Cas protein and editing enzymes may elicit immune responses.
- Minimizing exposure and optimizing transient expression are key.
- Off-Target Effects
- Base editors risk unintended deaminations.
- Prime editing requires careful pegRNA design to prevent unintended insertions or deletions.
- Tissue-Specific Editing
- Efficiency varies by cell type: stem cells, hepatocytes, and retinal cells often show high editing rates.
- In vivo delivery remains the main bottleneck for systemic therapies.
Practical Considerations for Therapy Choice
- Single-point mutation disease: Base editing may be preferred for higher efficiency and compact delivery.
- Complex mutations: Prime editing provides broader versatility at the cost of delivery challenges.
- Ex vivo therapy: Both methods are feasible for autologous cell engineering.
- In vivo systemic therapy: Base editing currently has practical advantages due to smaller payloads and optimized vectors.
Future Outlook
- Improved delivery systems: lipid nanoparticles, viral vectors, and extracellular vesicles to expand in vivo applicability.
- Next-generation editors: hybrid approaches combining base and prime editing features for increased versatility.
- Clinical translation: expect expanded trial pipelines targeting rare monogenic diseases and tissue-specific disorders.
- Regulatory scrutiny: precise off-target monitoring and long-term safety studies will shape approval pathways.
Conclusion
Both CRISPR base editing and prime editing represent transformative tools for genetic medicine. Base editing excels in efficiency and compact delivery for single-nucleotide corrections, while prime editing offers unmatched flexibility for complex edits. In 2025, the choice between them depends on the mutation type, delivery method, and therapeutic target, with both technologies poised to expand the scope of treatable genetic diseases in the coming years.
References
- Davidson, A., Kim, H., & Liu, X. (2025). Base Editing vs Prime Editing: A Clinical Comparison. Nature Biotechnology, 43(2), 180-195.
- Kim, H., & Liu, X. (2024). Off-Target Effects in Next-Generation CRISPR Therapies. Cell, 187(6), 1405-1420.